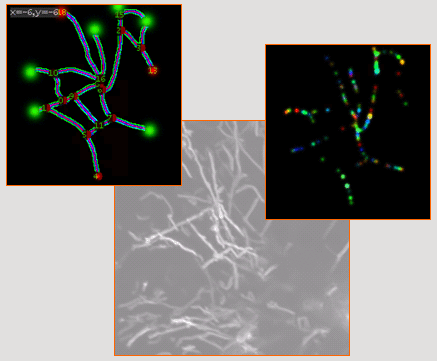
In this proposal, we plan to define new
methods dedicated to the analysis of nD microscopy data and to the
modeling of molecular and macromolecular mechanisms at the cell level.
In combination with the amount of information provided by
high-throughput nD microscopy
and automated image analysis methods (object segmentation, tracking,
...), we aim at designing computational and statistical models to
understand membrane trafficking and, more precisely to better elucidate
the role of Rab family proteins inside their multiprotein complexes.
Our main objective is then to provide computational methods and
mathematical models to automatically extract, organize and model
dynamic information observed in temporal series of images in
multi-dimensional (nD) microscopy. This represents a real scientific
challenge in applied mathematics, computer science and biology.
Nevertheless and beyond, we believe that
our studies will help to describe and better define the dynamic
architecture of multi-protein complexes involved in the time and space
regulation of many essential molecular mechanisms of the living, such
as the dynamic organization of the cytoskeleton, cytokinesis, nuclear
envelop biogenesis, cell adhesion,...
document
(pdf)