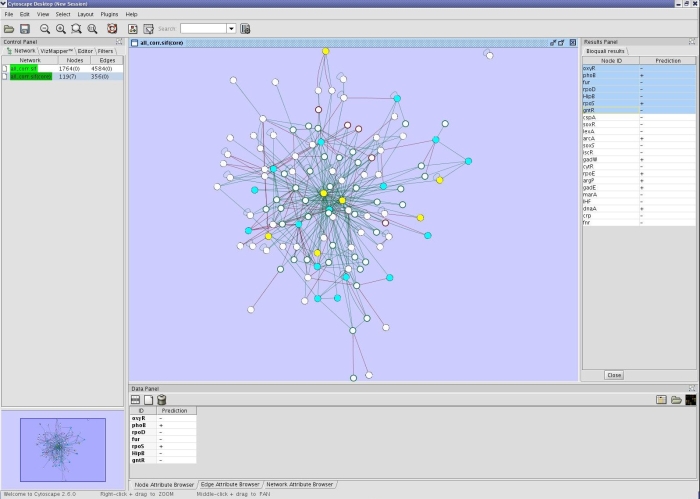
Description
BioQuali is a Plugin for Cytoscape developed at the Symbiose project at IRISA-INRIA, Rennes, France. The technical development and the integration of the Plugin has been carried on by the GenOuest Bioinformatics Platform.
BioQuali analyses regulatory networks and expression datasets by checking a global consistency between the regulatory model and the expression data. It diagnoses a regulatory network searching for the regulations that are not consistent with the expression data, and it outputs a set of genes which predicted expression is decided in order to explain the expression data provided.
The consistency of a network is understood in the following sense: "Each positive or negative variation of a node in the network, must be explained by a positive or negative total influence received from its predecessors". We have discretized this analysis by imposing that all nodes in the network are limited to be up or down regulated, and that the edges in the network can represent positive, negative, or unknown influences. However, for analyzing a global consistency all the nodes in the network must be taken into account. The BioQuali Cytoscape Plugin proposes the user to visualize the solution of this problem automatically and in few minutes no matter the size of the network.
BioQuali plugin bases all its functionalities in a Python library BioQuali available on line at (www.irisa.fr/symbiose/bioquali). As a Cytoscape plugin, it provides the visualization of this analysis in order to provide a more friendly environment and to encourage users of different disciplines to analyze their regulatory networks.